AAV serve as a flexible and economical tool for delivering genetic payloads required to carry out gene editing via CRISPR technology. PackGene provides customized vector cloning services for AAV-based CRISPR gene editing. Our AAV-CRISPR vectors for gene knockout or gene activation are divided into 5 categories, these categories are:
- SpCas9 and SpCas9HF available in our A and B series vectors
- SaCas9 and SaCas9HF available in our C and D series vectors
- AAV-CRISPR gene activation available in our E series vectors
- NmCas9 available in our F series vectors
- AsCpf1 and LbCpf1 available in our G, and H series vectors
CRISPR is a state-of-the-art genetic modification tool that can be used to knockout, mutate, or overexpress a gene of interest. CRISPR based research strategies have been widely employed across biological fields of study due to their powerful performance and relatively straightforward mechanism of action.
CRISPR gene editing requires three components: (1) a guide RNA (gRNA), (2) a Cas9 endonuclease, and (3) a protospacer adjacent motif (PAM) within the host genome. The gRNA are small single stranded RNA molecules that hold two distinct regions. The first gRNA region is a gene targeting sequence that is designed to complement a 15-24 base pair segment of the gene of interest’s DNA sequence and the second is gRNA component is a scaffold region. The gRNA scaffold region binds to a Cas9 endonuclease while leaving the host DNA compliment region of the gRNA unbound. The Cas9-gRNA complex may then scan the host cell’s genomic DNA for a specific 2-6 base pair sequence referred to as the PAM. Once the Cas9-gRNA complex recognizes the PAM sequence the gene targeting region of the gRNA may then anneal to the host DNA adjacent to the PAM sequence. After successful binding of the gRNA gene targeting region Cas9 will cleave the host DNA adjacent to the PAM sequence.
Cas9 mediated DNA cleavage generally results in gross disruptions in gene transcription and functional gene knockout following DNA repair. Alternatively, cleavage can be used to introduce precise DNA mutations or to activate gene expression using a modified gRNA scaffold region that inhibits Cas9 cleavage capabilities while simultaneously recruiting transcriptional proteins to a gene’s promoter region. Importantly, the specificity of target DNA recognition makes it feasible to design CRISPR based gene editing strategies targeted at nearly any gene.
The design and production of AAV vectors for CRISPR based gene editing can be both challenging and time-consuming. However, our professional team has experience and knowledge necessary to deliver high quality CRISPR-AAVs that are ready to use for your experimental needs immediately on delivery.
Ease of Use
PackGene maintains large database of pre-designed sgRNA sequences as well as of a library of reliable vectors. Simply pick out or upload your sgRNA sequence and select a vector from our library.
Expertise
PackGene’s experts have more than 10 years’ experience in the design, construction, and production of AAVs.
Efficiency
PackGene depth of expertise paired with our proprietary methods and optimized workflow ensure that creation of your vectors will move efficiently through design, production, and delivery.
PackGene’s proprietary SpCas9 expression vectors have been carefully designed to co-express SpCas9 and sgRNA from a single vector. First, in order to ensure that the SpCas9 expression sequence does not exceed the AAV vector insert length limit of ~5Kb we use the short but efficient promoter sequences EF1s or miniCMV. Secondly, we drive sgRNA expression using the efficient but relatively short H1 promoter to reduce to total bp length of the sgRNA expression cassette. With this approach we generated a single AAV vector capable of SpCas9 and sgRNA co-expression. Co-expression of SpCas9 and sgRNA from a single vector has many benefits over a classical duel vector approach since all infected cells express both components. This ultimately reduces the probability of gathering spurious data due to mixed cell expression phenotypes.
Our CRISPR vectors feature an optimized sgRNA scaffold that increases the cleavage efficiency of SpCas9HF by approximately 40%.
Our XA01 CRISPR vector is capable of SpCas9 and sgRNA co-expression from a single vector. PackGen’s XA01 uses a unique combination of short but efficient promoter sequences to drive expression of SpCas9 and sgRNA. This allows for co-expression of both essential components from a single vector.
Our XA03 through XA27 CRISPR vectors provide an option for sgRNA expression to be paired with various reporters. PackaGene’s XA03 to XA27 vectors hold U6-sgRNA expression cassettes with EFS, EF1, or CAG promoter-driven reporters including: mCherry, Cre, mCherry-2A-Cre, RenillaLuciferase-2A-Cre, ZsGreen, EGFP, EGFP-KASH.
Our XA02 CRISPR vector provides an option for SpCas9 expression without sgRNA. PackGene’s XA02 vector is a EFS promoter driven SpCas9 vector that does not contain sgRNA expression cassette.
We offer traditional SpCas9 expression vectors as well as high fidelity SpCas9HF expression vectors. SpCas9HF is a high fidelity variant of SpCas9 with very low off-target activity and similar on-target cleavage capabilities reletive to SpCas9 .
SaCas9 is a shorter version of Cas9 that holds similar cleavage activity to SpCas9 but is encoded by ~3.2kb gene versus the ~4.2kb of SpCas9. In addition to this reduced size the SaCas9 PAM (NNGRRT) is less commonly observed in host genomes at a frequency of approximately once in every 32bp, and the SaCas9 21nt-23nt target sequence is longer than that of traditional SpCas9. This decrease in PAM frequency and increase in target sequence length combine to result in higher SaCas9 binding specificity, and therefore a theoretically lower off-target cleavage rate than traditional SpCas9.
Increased cleavage efficiency through the use of an optimized sgRNA scaffold.
Reduced theoretical off-target cleavage rate.
SaCas9 and sgRNA expression in a single vector. PackGene’s C1 and D1 vectors hold SaCas9 or the high fidelity SaCas9HF expressed on a CMV promoter with sgRNA expression cassette integrated into a single vector.
The option for SaCas9 expression without sgRNA. Several PackGene vectors (C2, D2, C2XX, D2XX) are available for expression of SaCas9 or SaCas9HF without an integrated sgRNA expression cassette.
While Cas9 is commonly paired with traditional gRNA to produce targeted gene knockouts, Cas9 may alternatively be paired with a gRNA that holds a modified MS2 RNA linker scaffold. When paired with this modified gRNA, spCas9 is loses its DNA cleavage capabilities and gains the ability recruit the MS2-P65-HSF1 transcriptional activation component to drive downstream gene transcription. The net effect of this process is transcriptional activation at levels that can reach more than 1000-fold endogenous expression.
The 3.3-kb-long NmCas9 is an alternative version of Cas9 with similar activity to SpCas9. NmCas9 can be beneficial for applications where the theoretical off-target cleavage rate must be exceptionally low. Due to the recognition of the longer PAM (NNNNGATT), the frequency of the PAM sequence appears around once every 128 bp. In addition, the long target sequence length of 21-23nt provides an additional increase in target specificity. These features make the theoretical off target cleavage rate for NmCas9 lower than other Cas9 variants.
Easy Access
Reliable database and library, stored with large variety of vectors, simply upload your sgRNA sequence and pick the vector from our library.
Expertise
To ensure the quality and maximize the use of our products, PackGene recommends selecting 2-3 targets for non-validated gRNA.
Time Saving
With our experts specilized in this area with more than 10 years’ experience, we are confident that PackGene can carry out target design, map and vector production efficiently.
While Cas9 and its variants are the most commonly used CRISPR effector endonucleases, the Cpf1 endonuclease has gained increasing popularity as an alternative effector endonuclease for CRISPR gene editing. Like Cas9, Cpf1 requires a gRNA for activation, binds genomic DNA adjacent to a PAM sequence, and cleaves DNA following sgRNA target sequence annealing. Cpf1 is, however, is shorter in size than Cas9, and requires a shorter sgRNA for full functionality. Importantly, Cas9 cleavage results in blunt ended DNA while cleavage by Cpf1 results in 4-5base protruding sticky-ended DNA that offers a means for low-error and controllable gene insertions.
Cpf1 cleavage results in 4-5bp sticky DNA ends that are more amenable togenome modification applications.
We offer vectors for expression of AsCpf1 or LbCpf1.
- Products: CRISPR AAV vectors from PackGene Biotech
- Injection Method: Intramuscular Injection
- Recommended Dose: 25 μL, 1.5E+12 vg/ml
- Targeting Site: muscle
- Animal Model: DMD mouse model
- Journal: Genome Medicine, 2021 (IF=15.266)
- Paper Title: In vivo genome editing in mouse restores dystrophin expression in Duchenne muscular dystrophy patient muscle fibers
- DOI: 10.1186/s13073-021-00876-0
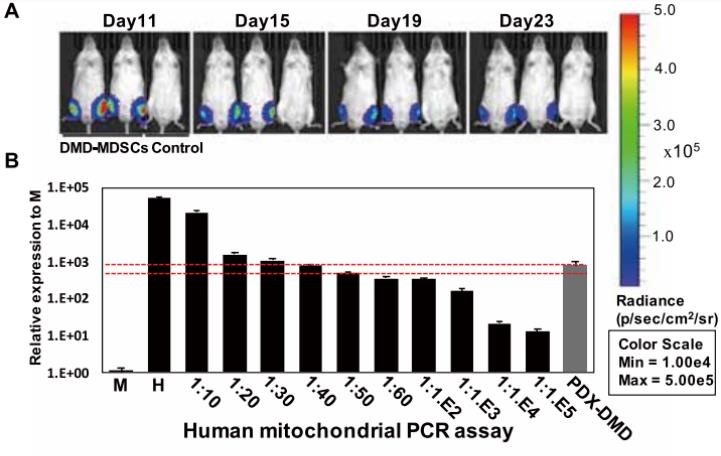
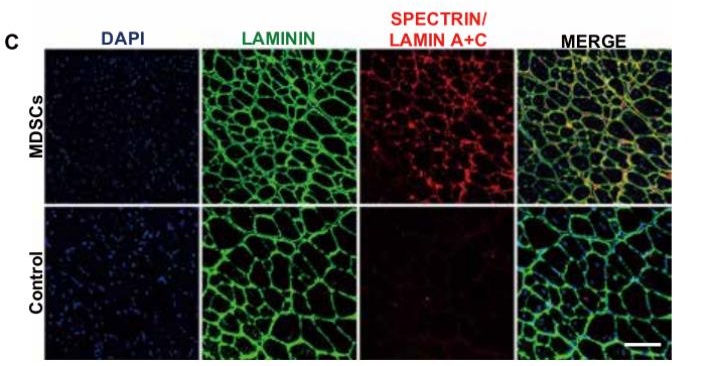
CRISPR AAV vectors were generated by PackGene Biotech Co.
Related Services / Products

Off the Shelf AAV Products
We offer a wide catalogue of pre-made AAVs for a multitude of research needs.
READ MORE

AAV Packaging Services
Ranging from pilot to industrial-scale AAV packaging for both in vitro and in vivo studies.
READ MORE

AAV Analytical Services
We are one of the few vendors that can do ALL in-house.
READ MORE
