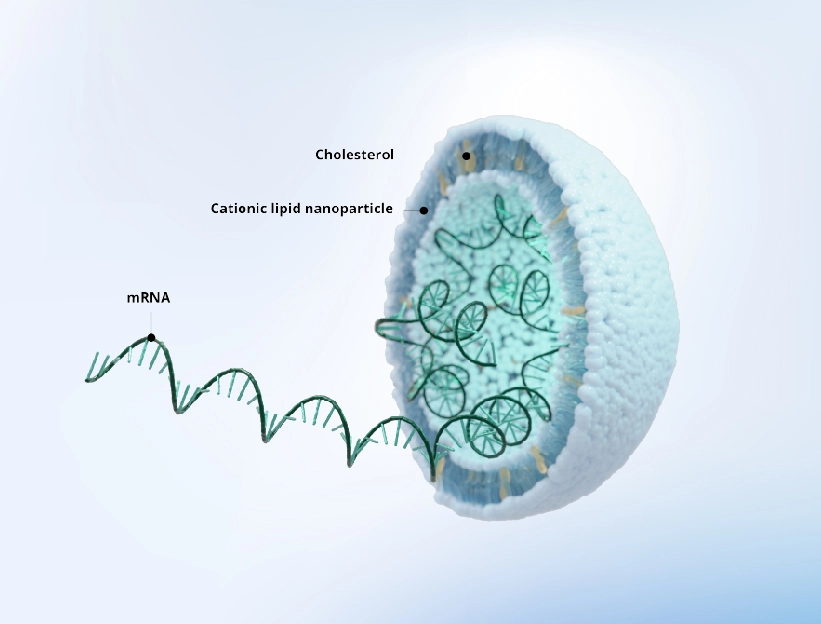
Key Benefits
-
Superior Quality
High mRNA integrity with a comprehensive QC panel -
Customizable Solutions
Flexible scales, enhanced stability, tailored modifications -
End-to-End Support
Expert guidance and scalable manufacturing
Service Details
| Catalog | Capping and modification | Gene | Quantity | Timeline(Business days) |
|---|---|---|---|---|
| mRNA | Custom mRNA Cap1 | Custom gene | 100μg-20mg | 10-15 |
| Custom mRNA Cap1 N1meΨU |
*Additional time (~2-3 weeks ) for custom gene synthesis
mRNA grade and QC standard
| QC Category | QC Item | Method | Specification | Research Grade | Add-on |
|---|---|---|---|---|---|
| Identification | Appearance | Visual Inspection | Clear and free of foreign particles | √ | |
| mRNA Concentration | UV Absorbance by Nanodrop | ≥1mg/ml | √ | ||
| RNA Integrity/size | Capillary Electrophoresis | Target ±30%, Expected band size detected | √ | ||
| Buffer Specification | Client Spec | RnaseFree H2O(default), PBS, 1mM Sodium Citrate, pH 6.4 | √ | ||
| Purity | A 260/280 Ratio | UV Absorbance by Nanodrop | 1.70-2.30 | √ | |
| Size based purity | Capillary Electrophoresis | >80% | √ | ||
| Impurity | Total Protein residue | Nano Orange | ≤1% | √ | |
| Plamsid DNA residue | qPCR | ≤0.1% | √ | ||
| dsRNA | Slot-blot | ≤1% | √ | ||
| Safety | Endotoxin | Semiquantitative LAL | <10EU/mg | √ | |
| Endotoxin | Quantitative | <10EU/mg | √ | ||
| Bioburden | Direct Inoculation | No growth after 48 hrs | √ | ||
| Raw material-Linearization | Linearization percentage | AGE | >95% | √ | HPLC |
| Host cell DNA | Quantitative PCR | ≤ 5% | √ | ||
| Total protein | Nano Orange | ≤1% | √ | ||
| RNA | AGE | Non-detectable by gel electrophoresis at 200ng | √ | ||
| Raw material -circular Plasmid | Gene of Interest | Sanger sequencing | 100% math reference sequence (not including poly A) | √ | |
| Poly A | Sanger sequencing | ≤110A ±5nt, ≤111-125A ±8nt | √ | ||
| Poly A Length | Enzyme digestion and CE | Target ±5% | √ |
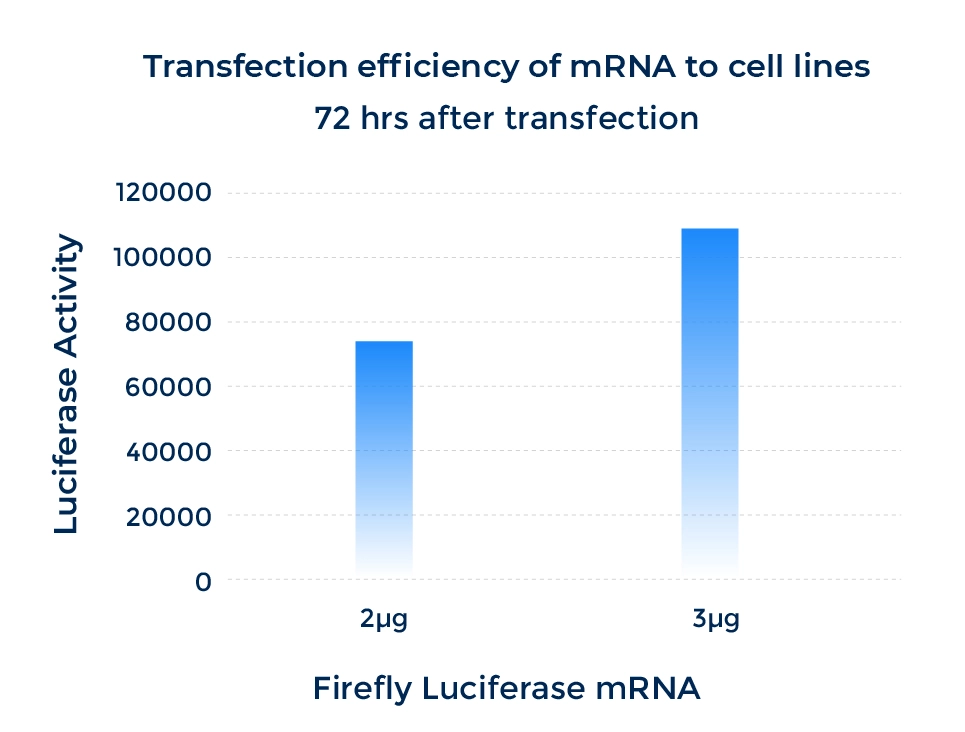
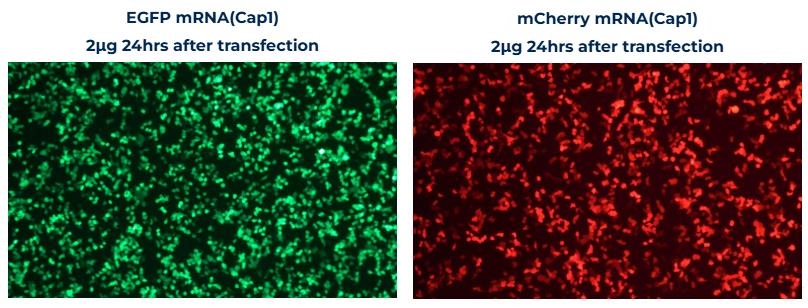
Performance
-
mRNA Integrity
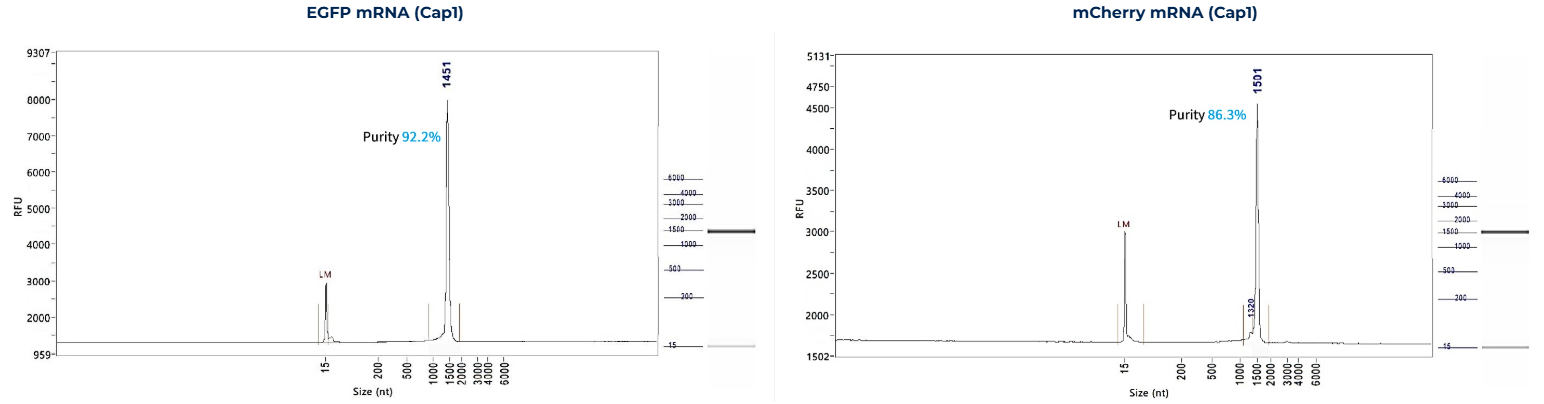

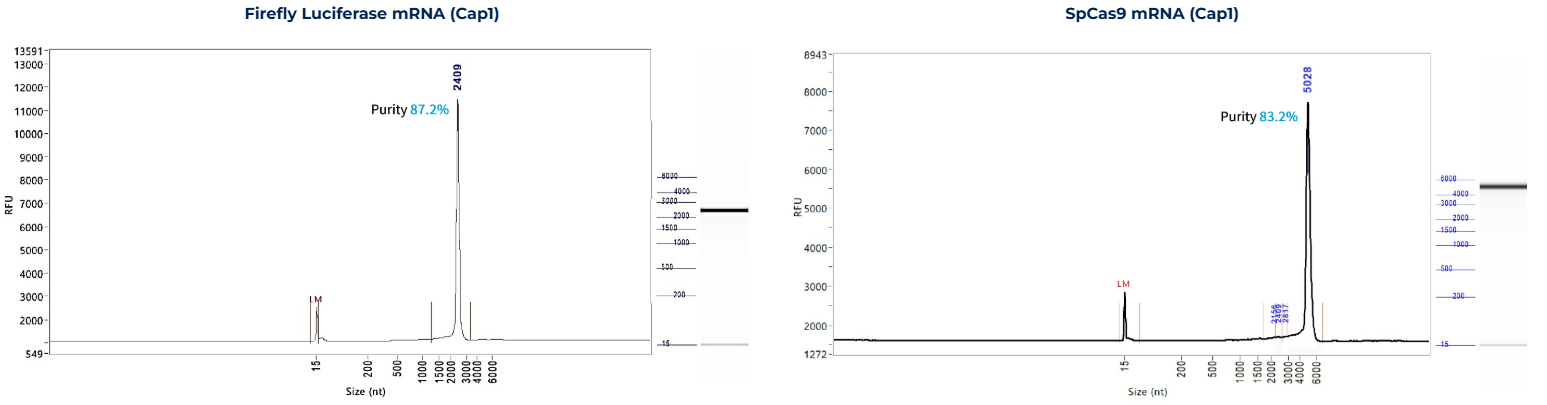
-
mRNA Expression